Smart Metabolites Database™ Ver.2 - Features
Smart Metabolites Database for GC-MS(/MS) Analysis
Automated, Highly Sensitive Detection of Metabolic Components by MRM Measurement
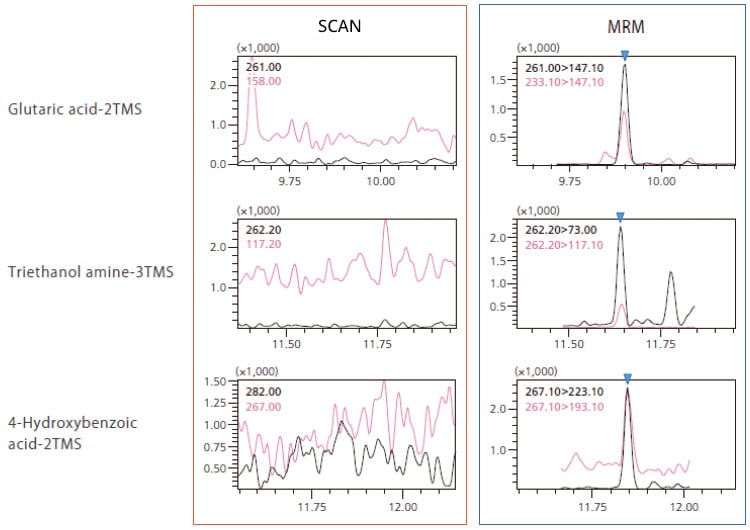
Metabolite analysis has conventionally been performed using scan methods; however, these methods often suffer from issues such as insufficient sensitivity to detect trace compounds, overlapping metabolites, and contaminants interfering with accurate results.
In this database, wide-target analysis using MRM can detect metabolic compounds with high sensitivity and selectivity, thus accurate results can be obtained even when analyzing for trace compounds. When combined with highly reliable metabolite identification based on retention indices, users can obtain highly accurate results in comparative analysis of foods and marker discovery.
Time and Labor Savings
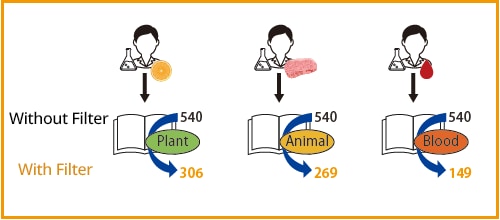
The Smart Metabolites Database includes more than 600 compounds and facilitates metabolite analysis across a wide variety of samples. Nevertheless, the greater the number of compounds analyzed, the more time needed for data analysis. To minimize this issue, the Smart Metabolites Database comes with a new filter function that allows users to select and analyze only those metabolic compounds likely to be detected in a given sample, thereby saving time and labor in the analysis process. This filter function can be turned on and off at user discretion.
-
Filter No. of Components All registered components (MRM) 540 Plant (food) 306 Animal (food) 269 Blood (plasma) 149 Urine 171 Cells 224 -
Offers Quantitative Sugar Analysis
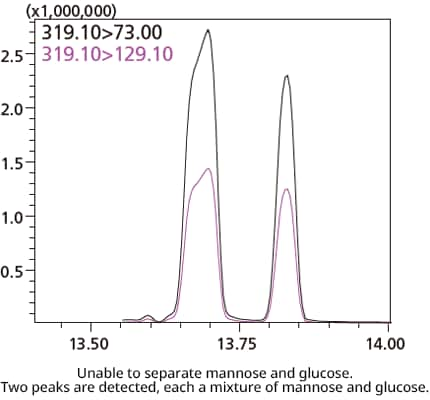
The methoxime-TMS derivatization method is in common use, but produces multiple geometric isomers from reducing sugars and causes chain formation due to derivatization, making chromatographic separation of sugar isomers not possible.
A new sugar quantification method uses an optimized derivatization method to enable the selective detection of sugars. The database also contains compound information and calibration curve information needed to analyze 24 major sugars, allowing semi-quantitative results to be calculated for detected sugars.
The extraction procedure used on samples is identical to that used by simultaneous metabolite analysis, thus simultaneous metabolite analysis and quantitative sugar analysis can both be performed on the same processed sample.
-
Methoxime-TMS Method
-
Sugar-Specific Method
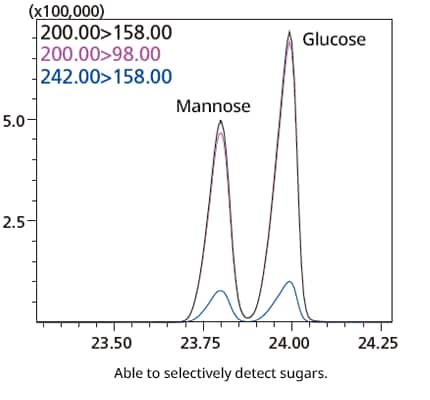
*Semi-quantitative results may vary significantly from true values depending on equipment conditions and sample preparation. Quantitative tests should be performed with standard samples if accurate quantitative results are required.
Total Support for Food Metabolomics Analysis
Shimadzu provides complete support for GC/MS food metabolomics analysis from sample preparation to visualization on metabolic maps and multivariate analysis. The "Metabolomics Handbook" compiles practical information on food sample preparation, allowing even newcomers to food metabolomics to output results up to and including multivariate analysis.
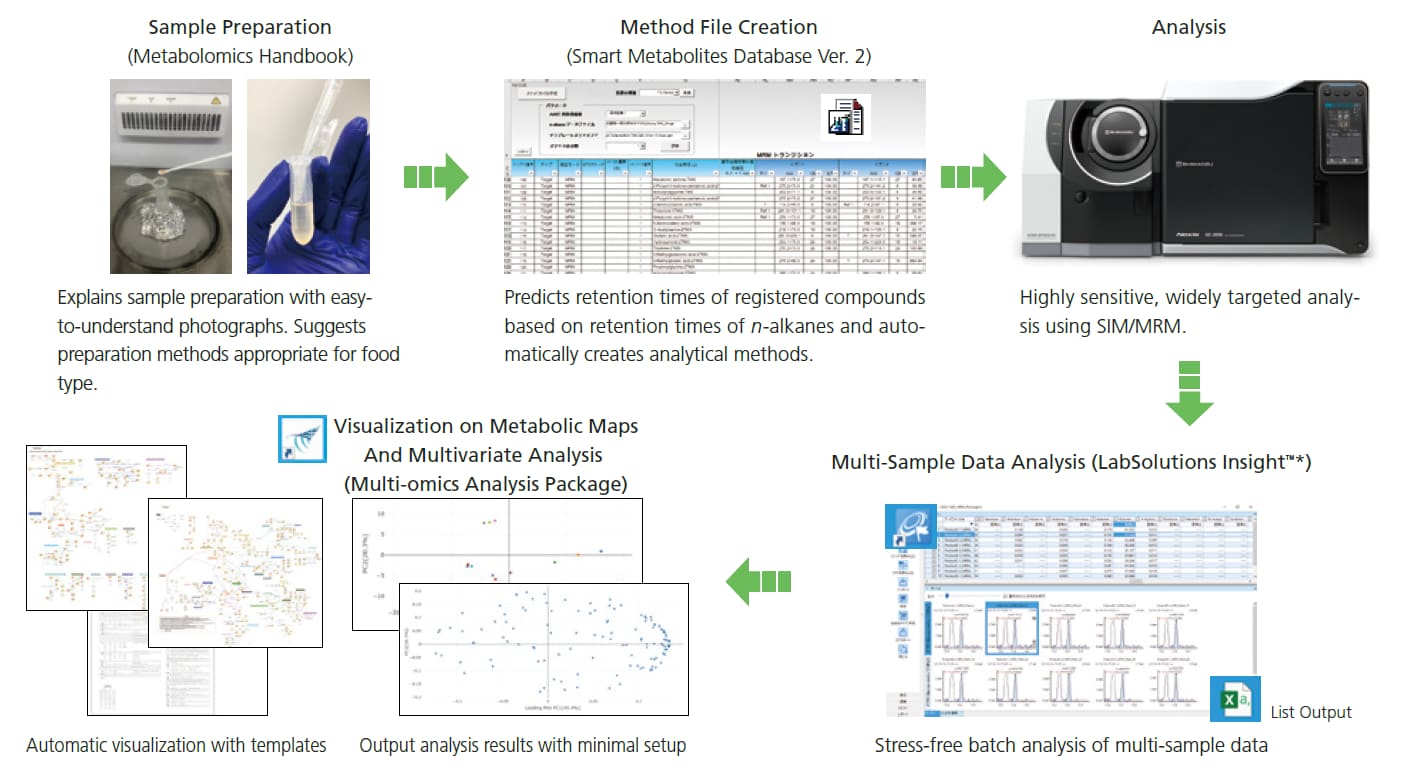
* LabSolutions Insight and Multi-omics Analysis Package are not included in this product.


