LC/MS/MS MRM Library for Phospholipid Profiling
For LabSolutions™ LCMS

This MRM library includes two methods: one for phospholipid classification by comprehensive analysis of the main phospholipids in biological samples, and one for fatty acid composition determination created using analytical results obtained with the classification method. The library targets phospholipids containing C14 to C22 fatty acids, and includes MRM transitions for up to 867 components. This library enables performing phospholipid profiling by conducting an initial analysis with a phospholipid classification method. This is followed by creating a method for fatty acid composition determination based on the phospholipid peak detected in the first analysis, and subsequently using this method to perform a second analysis to determine fatty acid composition.
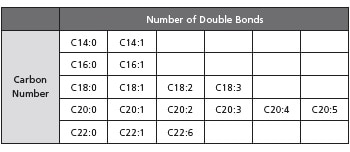
Target Phospholipids
The phospholipids registered in the MRM library have fatty acid compositions with the carbon number and double bond combinations shown in the table below. The phospholipid targets of the library are phosphatidylcholines (PC) with a lyso-group, phosphatidylethanolamines (PE), phosphatidylglycerol (PG), phosphatidylinositol (PI), phosphatidylserines (PS), and sphingomyelins (SM).
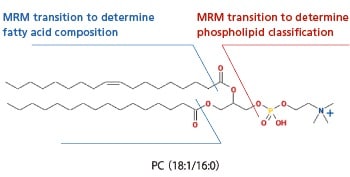
MRM Transition Composition
Focusing on the characteristic phospholipid head group, the library includes a method for phospholipid classification and a method for fatty acid composition determination (for fatty acid compositions of the given combinations) that make use of these MRM transitions. The figure below shows each MRM transition required to identify PC (18:1/16:0). The phospholipid can be inferred by combining the analytical results obtained from these MRM transitions.
Step 1 Simultaneous Analysis Using the Phospholipid Classification Method
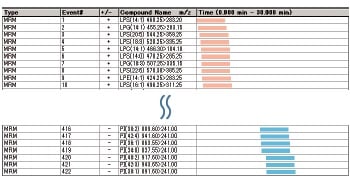
The phospholipid classification method is capable of comprehensive analysis of the main phospholipids in biological samples. It determines the phospholipid class based on the main characteristic head groups of phospholipids. Its analysis targets are phospholipids that include C14 to C22 fatty acids.
Step 2 Peak Identification
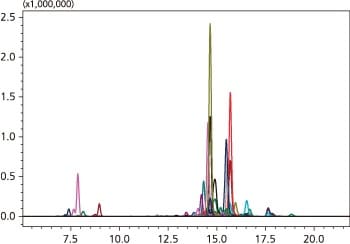
The phospholipid peak detected by the phospholipid classification method is identified. The structural information on the detected phospholipid at this point is the phospholipid class (PC, PE, PG, PI, PS, SM), and the total carbon number and number of double bonds in its constituent fatty acids. In the next step, the fatty acid composition determination method is created.
Step 3 MRM Event-Link Editor Editing the Fatty Acid Composition Determination Method
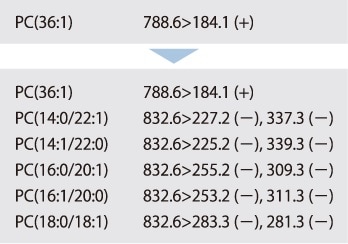
Software allows a fatty acid composition determination method to be edited from 867 MRM transitions for the phospholipid peak detected during first analysis.
Step 4 Simultaneous Analysis by Fatty Acid Composition Determination Method

Edited in MRM Event-Link Editor, the fatty acid composition determination method is used to perform a second analysis on the same sample. Phospholipid profiling can be performed based on the results of the first and second analysis.
Step 5 Data Analysis
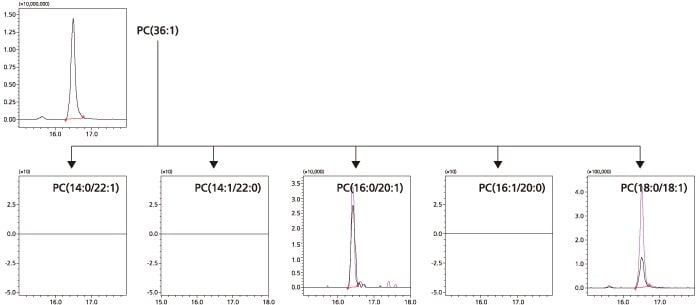
Based on the analysis results, it was determined that the sample contains PC (16:0/20:1) and PC (18:0/18:1), as shown in the figures below.
Remarks and Precautions
LabSolutions LCMS Ver. 5.109 or later and LabSolutions Insight Ver. 3.8SP1 or later are required.
LabSolutions is a trademark of Shimadzu Corporation or its affiliated companies in Japan and/or other countries.
For Research Use Only. Not for use in diagnostic procedures.
News / Events
-
LabSolutions Insight Profiler, Non-Targeted Analysis Software for LC/QTOF Data has been released
LabSolutions Insight Profiler is a unified, end-to-end workflow for LC/QTOF data processing — with a "single-click" approach. Insight Profiler is purpose-built for non-targeted analysis in suspect screening, unknown identification, and metabolomics across complex sample sets.
-
Cell Culture Profiling for LCMS-2050
This package includes a preconfigured method covering vitamins, amino acids, and other culture media components and metabolites secreted by cells.
-
Shimadzu has released the LCMS-8065XE
The new LCMS-8065XE is a triple-quadrupole mass spectrometer with EVOLVED, EFFICIENT, and EXACT capabilities. These exceptional capabilities ensure high reliability and enhanced productivity, empowering the laboratory for the future.
-
Latest issue of Shimadzu Journal, featuring Environmental Analysis, has come out.
This issue showcases advanced technologies and research tackling the global challenges posed by PFAS. As part of Shimadzu’s ongoing commitment to sustainability and problem solving, we strive to reduce environmental impacts and build a better future.
-
New High Resolution Accurate Mass Library for Forensic Toxicology
Perform forensic toxicology screening for drugs of abuse, psychotropic drugs, pharmaceuticals, pesticides, and natural toxins using this high-resolution accurate mass database.
-
Shimadzu has released the Shim-vial™ H glass, S glass.
Shimadzu provides high-quality vials that thoroughly eliminate these risks by visually inspecting each vial, allowing them to be used with confidence.


