Nexera-e - Features
Comprehensive Two-Dimensional Liquid Chromatograph
Maximizing the full potential of two separation modes to perform comprehensive two-dimensional separation
Comprehensive 2D-LC is an analytical methodology combining two independent separation modes orthogonally, greatly increasing separation efficiency. The combination of different modes enables the separation of peaks that are difficult to separate using conventional LC, providing excellent results for the analysis of complex sample matrices.
 |
|||||
|
|
||||
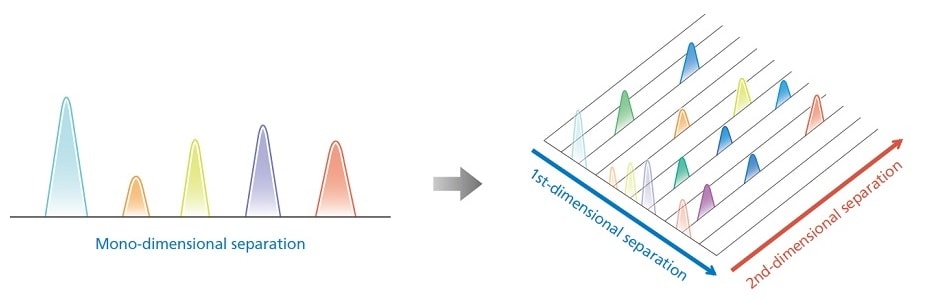
Differences between a conventional 1D-LC and a comprehensive 2D-LC
In comprehensive 2D-LC, all eluates from the first dimension can also be analyzed in the second dimension. The eluates from the first dimension are injected on to the column in the second dimension using two sampling loops that are alternately repeated by a switching valve.
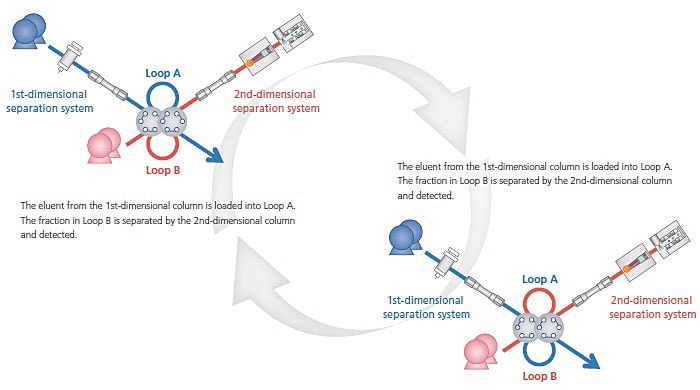
Flow line and mechanism of the comprehensive 2D-LC system
Simultaneous detection in positive and negative mode
Shimadzu triple quadrupole mass spectrometers (TQMS) have the world-class polarity reversal speed (5 ms) and data capture speed (1.5 ms). In comprehensive 2D-LC analysis, the analysis time of second dimension is short, therefore the high-speed performance of the LC-MS affects the quality of the data. Thanks to the high-speed analytical performance of the Shimadzu TQMS, it was possible to obtain stable data even for high-speed analysis of multiple component in comprehensive 2D-LC.
The figure below shows contour plotting data obtained from simultaneous analysis of 10 polyphenol components using Nexera-e with shimadzu TQMS. Although some components are detected in positive mode and others in negative mode, it is possible to acquire both data in a single analysis using the Shimadzu TQMS.
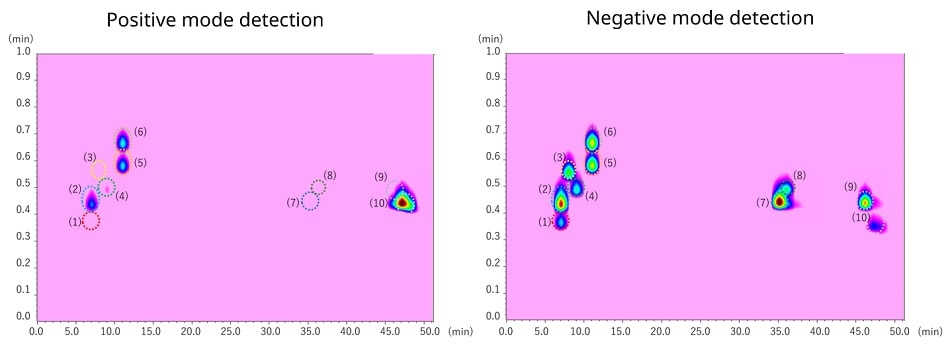
Simultaneous analysis of polyphenols
(1) gallic acid, (2) catechin, (3) hesperidin, (4) quercetin-3-o-glucoside, (5) isorhamnetin-3-o-glucoside,
(6) sinapic acid, (7) quercetin, (8) naringenin, (9) kaempferol, (10) apigenin
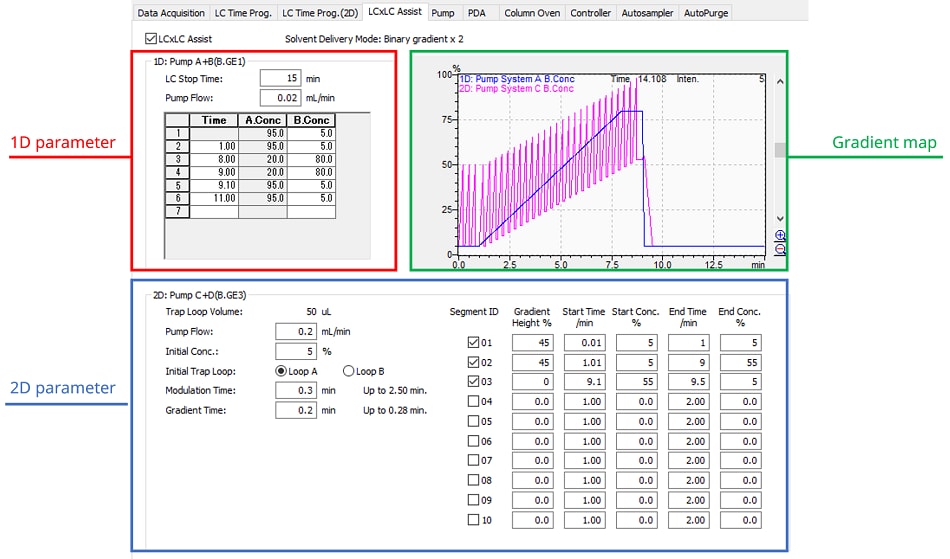
Easy setting of 1D and 2D analysis parameters

Comprehensive 2D-LC requires the setting of analytical parameters for the first and second dimensions, respectively. However, setting up the First dimension for ultra-low flow rate analysis and the second dimension for ultra-high-speed analysis can be complicated.
Nexera-e has a dedicated support function to facilitate these settings. The analytical conditions for the first dimension can be set up in the same way as for normal analytical LC, and the gradient settings for the second dimension are automatically generated based on the modulation time.
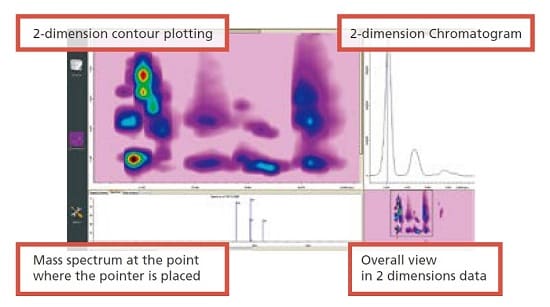
2D qualitative and quantitative analysis using Contour Graphics
Acquired data is converted to two-dimensional contour plotting using ChromSquare, the software for comprehensive 2D-LC analysis. A peak on a chromatogram is recognized as a spot on the contour plot. The qualitative and quantitative data processing are performed for the target spot.
-
-
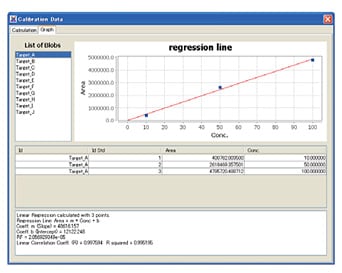
Calibration curve